Simulating Metabolism with Statistical Thermodynamics
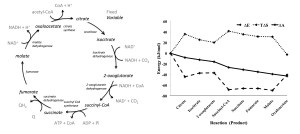
New methods are needed for large scale modeling of metabolism that predict metabolite levels and characterize the thermodynamics of individual reactions and pathways. Current approaches use either kinetic simulations, which are difficult to extend to large networks of reactions because of the need for rate constants, or flux-based methods, which have a large number of feasible solutions because they are unconstrained by the law of mass action. This report presents an alternative modeling approach based on statistical thermodynamics. The principles of this approach are demonstrated using a simple set of coupled reactions, and then the system is characterized with respect to the changes in energy, entropy, free energy, and entropy production. Finally, the physical and biochemical insights that this approach can provide for metabolism are demonstrated by application to the tricarboxylic acid (TCA) cycle of Escherichia coli. The reaction and pathway thermodynamics are evaluated and predictions are made regarding changes in concentration of TCA cycle intermediates due to 10- and 100-fold changes in the ratio of NAD+:NADH concentrations. Finally, the assumptions and caveats regarding the use of statistical thermodynamics to model non-equilibrium reactions are discussed.
See the complete article: http://t.co/7jvyGryvHr



